So you think you have a promising hit?
My mom has four boxes of turmeric tea sitting in her pantry. I suspect this has something to do with all of the articles and studies over the years purporting “turmeric may help heal ____." For an overview of why turmeric is probably not helping ____ much, see Derek Lowe’s great post on its most infamous ingredient Curcumin Will Waste Your Time.
The problem is that curcumin is a classic “nuisance compound” that will show up as a hit in many screening assays, and tempt researchers and funders away from solid science. A great new review Nuisance compounds in cellular assays by Dahlin and coworkers makes clear that curcumin is not an isolated example, but rather representative of a persistent problem in phenotypic drug discovery. Thus, a recent focus of us at PostEra Manifold is providing better ways for researchers to know if their hits have these known nuisance activities or promiscuous moieties.
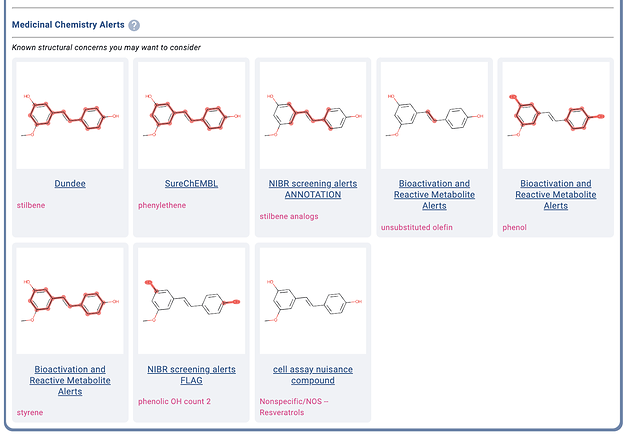
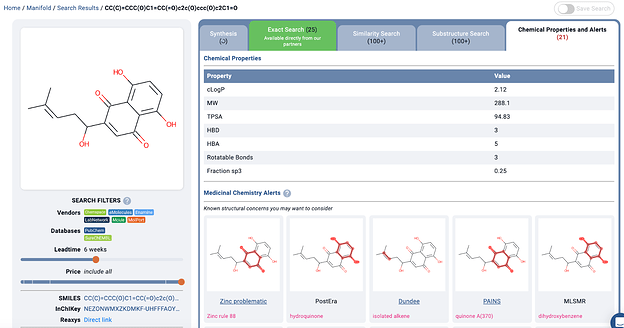
The Manifold med-chem alerts output for an example from another common class of nuisance compounds, Resveratrols. (You can also try curcumin in Manifold; it triggers so many alerts, it was hard to fit in the post
)
We believe users should be able to quickly check if their compounds are known to interfere with assay technologies or produce biological effects through undesirable modes of action. Knowing if a compound may be a suspected aggregator, redox cycler, CAD, fluorescent compound, reporter inhibitor, chelator, non-specific electrophile, etc. allows for improved counter-screening procedures, and thus higher quality hits and less false positives.
Manifold’s med-chem-alerts (also available via API) are now expanded to check for cell-free and cell-based assay nuisance behavior. The rest of this post will delve further into the need for these alerts and their origin.
An overview of difficulties in phenotypic hit validation and triage
Phenotypic screening, as opposed to target-based drug discovery efforts, focuses on first screening for biological readouts without pre-specifying a drug-target of interest. Nuisance compounds that "exhibit assay interference or undesirable mechanisms of bioactivity” [Dahlin et al.] are everywhere in phenotypic and high-content screening, and waste precious resources when wrongly pursued into more expensive stages of drug discovery.
To be clear, this problem is not isolated to cell-based assays, which are often quite specific to the disease area. False positives are rampant in cell-free assays of protein targets, and still pose huge issues for screening efforts. It’s just that it has become more clear how to deal with some of these issues (though it’s still hard and resource-consuming to do right!)
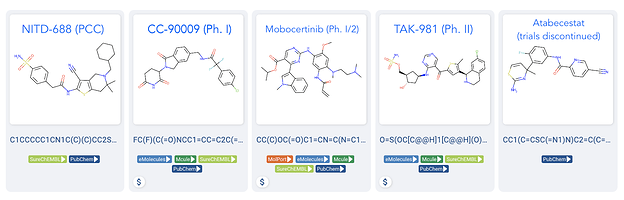
The recent resurgence and success of phenotypic drug discovery. Examples from drughunter.com’s Feb 2021 - Small Molecules of the Month of small molecules discovered through phenotypic or cellular screening efforts. Visit the original article to see how Takeda has even started a large number of programs after interrogating interesting mechanisms found from phenotypic hits.
As phenotypic drug discovery makes its impressive return, it has also become clear that nuisance compounds are not going away. One only has to look at all of the excellent recent articles around this topic to know it is of significant interest:
“key challenges need resolution: progression of poorly qualified leads and overloaded pipelines due to lack of effective tools to process and prioritize hits; and advancement of leads with undesirable mechanisms that fail at more expensive stages of discovery.” - The future of phenotypic drug discovery, Cell Chemical Biology 28, March 18, 2021
“In our view, a major obstacle to realizing the full potential of chemical biology will not just be difficult-to-drug targets or even the sheer number of targets, but rather nuisance compounds, due to their ability to waste significant resources and erode scientific trust.” - Nuisance compounds in cellular assays, Cell Chemical Biology. February 15, 2021
“Whereas this process [of hit triage and validation] is usually straightforward for target screening hits, phenotypic screening hits act through a variety of mostly unknown mechanisms within a large and poorly understood biological space” Hit Triage and Validation in Phenotypic Screening: Considerations and Strategies, Cell Chemical Biology. November 19, 2020
“These promises [of phenotypic screening] come with the inherent challenges of… a more complex hit-validation process, more complex structure-activity relationships and the unknown safety risks associated with the mode of action of the identified hits.” The European Federation for Medicinal Chemistry and Chemical Biology (EFMC) Best Practice Initiative: Phenotypic Drug Discovery ChemMedChem. April 07, 2021
At PostEra, we often help partners at the hit discovery and hit expansion stages. One challenge, in both target-based and phenotypic screens, when dealing with many compounds, is knowing if a molecule contains previously observed assay interference or promiscuous moieties, or is even a nuisance compound itself.
All of the above publications mention the usefulness of checking hits against alerts (such as PAINS, Eli Lilly’s alerts, etc.), but it is often hard to find the alerts all in one place. These lists of alerts exist and have been published across multiple Pharma companies, but it is time consuming to check molecules against all these separate lists.
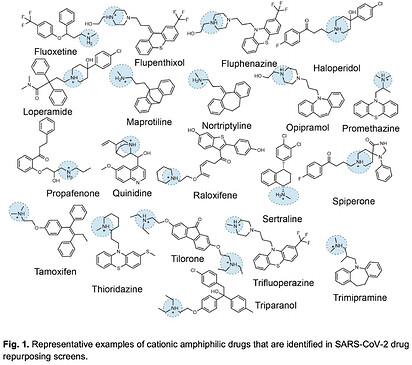
An example from COVID 19. A recent preprint Phospholipidosis is a shared mechanism underlying the in vitro antiviral activity of many repurposed drugs against SARS-CoV-2 showed that many of the repurposed drugs found to have SARS-CoV-2 antiviral activity were cationic amphiphilic drugs (CADs). These molecules are known to be confounders in cell-based screens due to their induction of phospholipidosis, which the authors showed was largely responsible for their antiviral activity.
Now, Manifold makes it easy for users to check a molecule for these problematic “med-chem alerts.” The list builds on many prior works including:
- Many of the alert sets mentioned by Pat Walters in his blog post on Filtering Chemical Libraries
- Other alerts from sets including those in recent papers such as Evolution of Novartis’ Small Molecule Screening Deck - Design and Eli Lilly’s alerts mentioned in Rules for Identifying Potentially Reactive or Promiscuous Compounds .
- In-house modifications to published alerts and custom written alerts for screening results.
- Bioactivation and reactive metabolite alerts form multiple published sources.
- Cell-based assay nuisance compounds from the aforementioned Nuisance compounds in cellular assays
Obviously, these alerts should be taken as guidance rather than gospel, and it should be recognized that each alert set was not developed for the exact same application. However, we’ve found that when dealing with a large number of hits, they are very useful for triaging those hits that should go on to more extensive experimental validation.
Furthermore, these alerts are certainly useful in designing screening sets for both target-based and cell-based assays. If one can excise the problematic structures early on, then they will not have to deal with them as confounding screening results later.
For cell-based assays, it’s also very helpful to design sets of both nuisance compounds, and well-validated chemical probes targeting known pathways. The biological readouts from these sets can then be compared to the readouts from the hits and inform future triage and mode of action investigation.
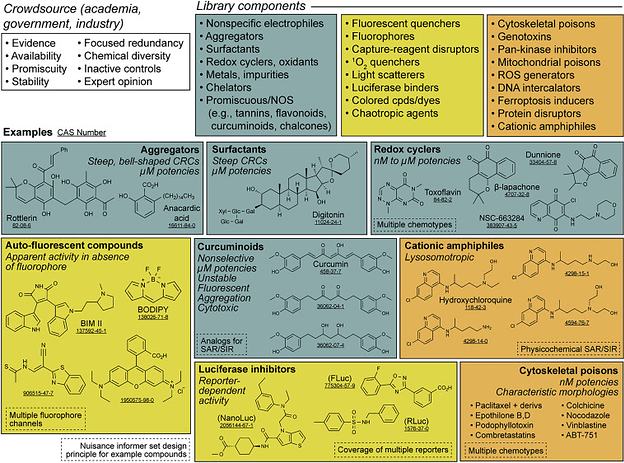
Many of the frequent nuisance compound classes one should look out for when running cell-based screens, from Nuisance compounds in cellular assays. The authors have compiled a large list of many of these problematic compounds, which is now searchable at https://postera.ai/manifold
Now that this set of med-chem-alerts and known nuisance compounds is easily searchable, hopefully users can triage hits effectively and understand which are suspected of interfering with assay technologies, and which may be working through undesirable modes of action. Getting all of this right early on is a lot better than realizing mistakes after the fact.
For ideas of how expensive these mistakes can be, estimates are that more than $150 million USD of NIH funding has been spent on curcumin, with no result. As a more recent example, this new paper detailing a classic class of nuisance compounds, cationic amphiphilic drugs (CADs), Phospholipidosis is a shared mechanism underlying the in vitro antiviral activity of many repurposed drugs against SARS-CoV-2. In that paper, the authors “estimate the cost of the clinical trials component alone, over the last year, for phospholipidosis-inducing CADs to be over $6 billion US dollars.”
Another example from COVID 19, in a cell-free assay. Shikonin is a component of chinese herbal medicines. It and similar naphthoquinones are known nuisance compounds. After reports of activity, a lot of effort was taken in the recent article Ebselen, Disulfiram, Carmofur, PX-12, Tideglusib, and Shikonin Are Nonspecific Promiscuous SARS-CoV-2 Main Protease Inhibitors to show that the inhibition of SARS-CoV-2 proteases was non-specific. Its weak density in a crystal structure then was noticed by a researcher looking into structures of ligands bound to MPro.
Now obviously, the ability to easily check compounds for medicinal chemistry alerts and potential nuisance behavior is not going to solve the issues of publication pressures or the need to quickly show results for a project. But making these filters more accessible will hopefully make the ubiquity of false positives and undesirable modes of action widely known. It’s likely too late for the turmeric tea in my mom’s cupboard, but here’s hoping we can save a lot of effort in the future.