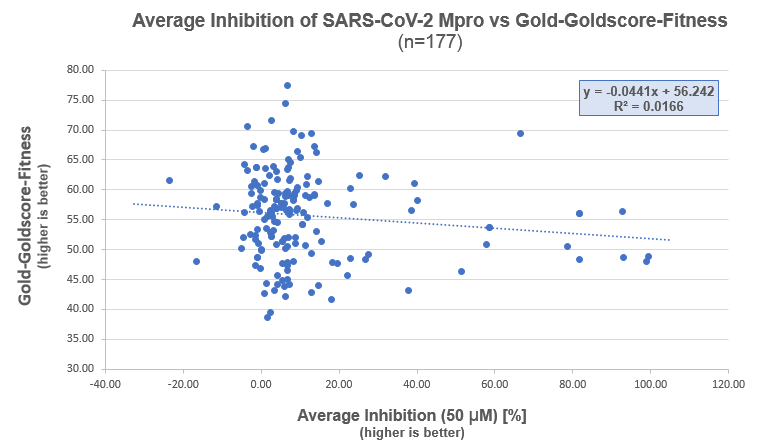
We’ve docked the non-covalent ligands available as of the 27th April from the CDD Vault using GOLD. GOLD was configured to create an ensemble of 6 proteins taken from the Fragment Screening results selected by diversity based on overall protein backbone RMSD. The top ranked pose by fitness was kept from each docking.
This led to ~3000 docked molecules.
We then re-ranked the molecules based on certain binding properties, effectively using a binary decision tree. In summary:
-
Structures with a fitness > 60.0 were ranked before structures with a fitness < 60.0
-
In these two sets, the structures were further ranked. Structures forming an H-bond to GLU166 and HIS163 were ranked highest, followed by structures that formed at least one H-Bond to one of these residues.
-
The resultant subsets were then further ranked based on whether the structures formed an H-bond to HIS41
-
The larger array of subsets was then ranked based on buriedness calculated using GOLDMine - structures more than 90% buried were ranked ahead of structures that were less than 90% buried
-
Finally, ties on the basis of the above were ordered by fitness.
The files are available at the dropbox link below:
The score based on the above has the tag . By way of example, a score might be
22120062.9252
This seems big, but its actually an indexing: the first 2 signifies “Scores more than 60.0” the second 2 is a sum of two 1s - the first 1 indicates ‘doesnt form an H-bond to GLU166’, the second indicates ‘doesnt form an H-bond to HIS163’.
The next 1 indicates that this doesn’t form an H-Bond to HIS41. The final ‘2’ indicates this is buried
Finally the fitness score is added. So we can read this as a highish scoring docking by fitness, that is buried but doesn’t seem to form certain key interactions in the binding site in the pose the program has produced.
The original GOLDScore is also included in the SD Files with the tag so sorting the results on this alone is possible if the user would rather interpret the data on this basis alone.
I note that none of the poses seem to form an H-Bond to all 3 of GLU166, HIS163 and HIS41. This is unlike some known inhibitors (for example 6Y2G) in the active pocket.